Q1 2026 Platform Update: Advancing Viral Surveillance, Sequencing Flexibility, AMR Prediction, and EPI2ME Integration¶
The landscape of clinical microbiology and public health is evolving faster than ever. At BugSeq, we are continuously enhancing our platform to ensure laboratories have the most advanced, accurate, and rapid bioinformatics tools at their disposal. In the first quarter of 2026, we focused our efforts on four major areas: deepening our viral surveillance capabilities, optimizing our pipelines for the rapid turnaround of the new Illumina MiSeq i100, expanding our expert-curated Antimicrobial Resistance (AMR) databases, and seamlessly integrating with Oxford Nanopore’s EPI2ME Desktop environment.
Recently, we also released BugID, a machine learning enabled tool to implement evidence-based reporting thresholds specific to your NGS assay; see the dedicated blog post for details.
These innovations empower our users to make faster, more confident decisions, whether tracking seasonal viruses, managing hospital outbreaks, or treating highly resistant infections.
Seamless BugSeq Integration with EPI2ME Desktop¶
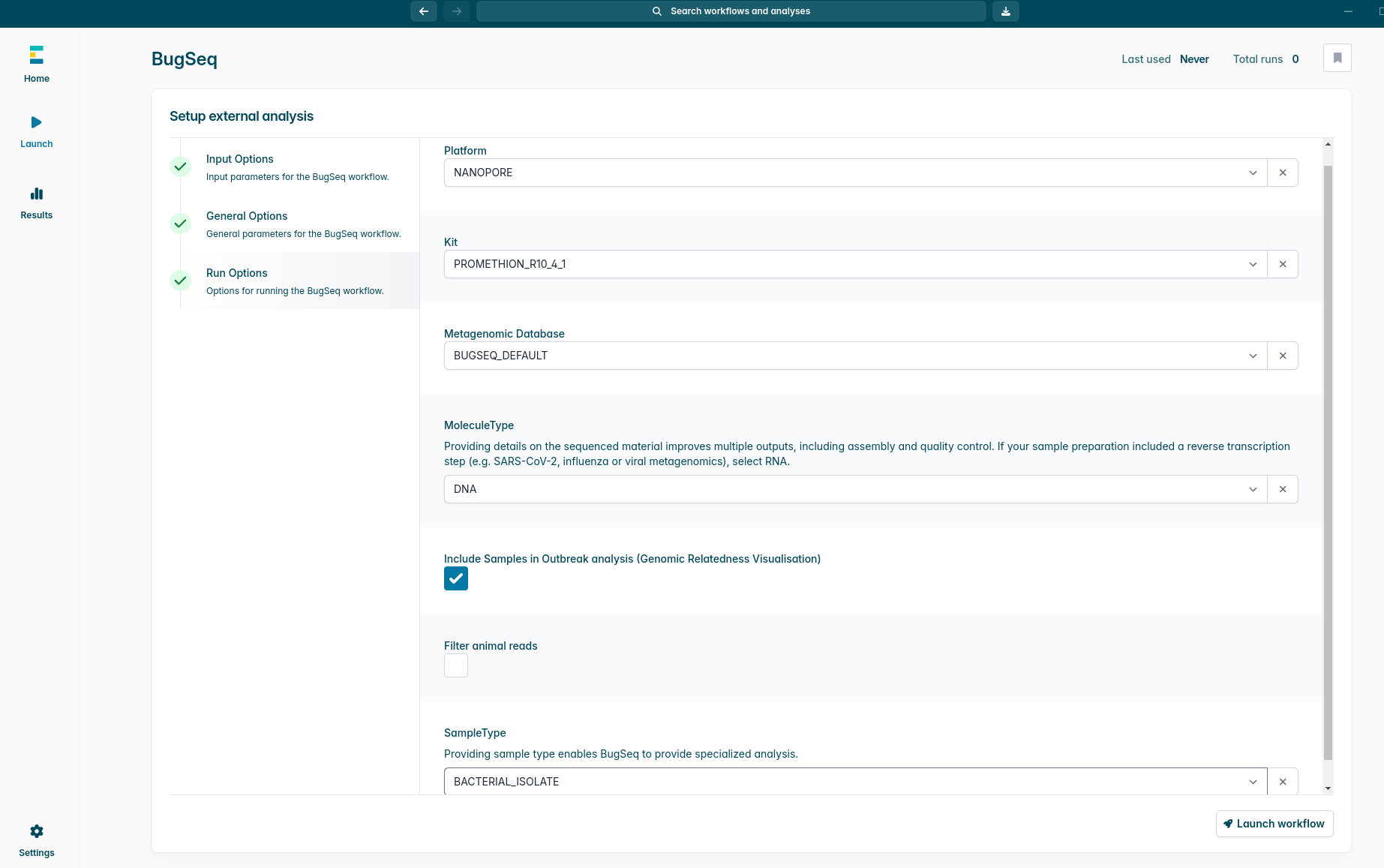
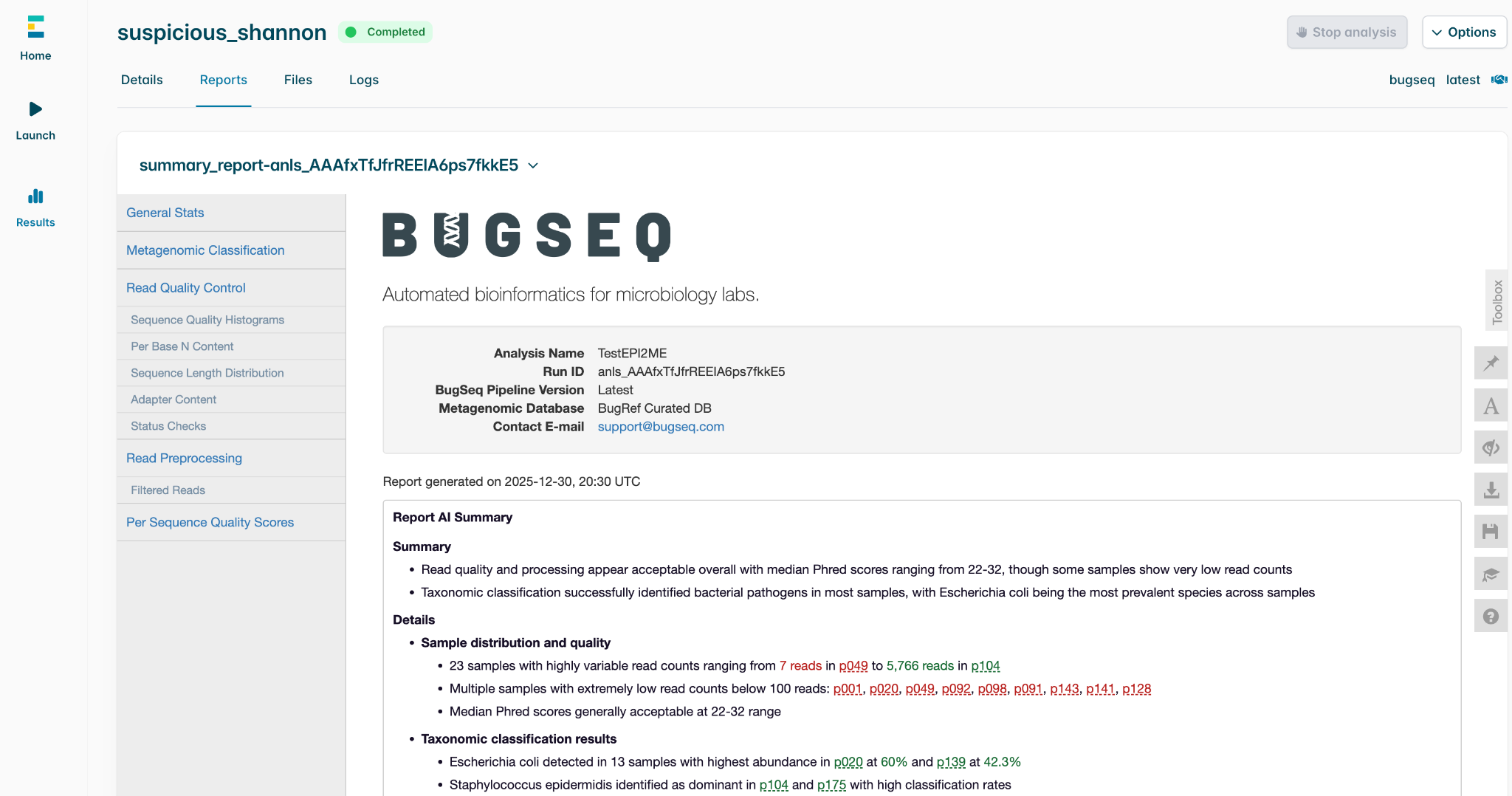
Nanopore sequencing provides unparalleled real-time insights, but moving data between different analysis ecosystems can cause workflow friction. To eliminate this bottleneck, we are thrilled to announce that BugSeq analysis is now officially integrated into the latest EPI2ME Desktop v5.4.0 release.
This direct integration allows users to submit supported Oxford Nanopore data to the BugSeq cloud platform seamlessly from within the EPI2ME Desktop interface. Once the BugSeq cloud completes the analysis, results are available directly within EPI2ME.
Advancing Viral Surveillance and Wastewater Monitoring¶
As viral threats continue to shift, high-resolution genomic surveillance remains our best defense. In Q1 2026, we rolled out major updates to our viral analysis pipelines:
- Subtype-specific influenza reference sequences: We have refined our influenza analysis to utilize subtype-specific reference sequences. This provides significantly higher accuracy for variant calling, consensus sequence generation and tracking specific flu lineages.
- Nextclade analysis for all Influenza A H/N types: The threat of a flu pandemic demands broad surveillance. We’ve expanded our Nextclade integration to support all Influenza A Hemagglutinin (H) and Neuraminidase (N) types with Nextclade databases available. This ensures public health labs can seamlessly analyze avian, swine, and other non-human influenza strains with the same ease as seasonal human flu, bolstering pandemic preparedness and early detection.
- Wastewater analysis for RSV: Respiratory Syncytial Virus (RSV) poses a severe annual burden on pediatric and elderly populations. We have incorporated RSV into our wastewater analysis pipeline to enable early subtype and lineage characterization.
Optimized Support for Illumina MiSeq i100 Data¶
The introduction of the Illumina MiSeq i100 has brought a new level of speed to short read sequencing. Its rapid run mode is a game-changer for time-sensitive applications like infection prevention and clinical metagenomics. To ensure our users get the most out of this instrument, we have optimized BugSeq specifically for i100 data:
- Better short-read single-end assembly: Rapid sequencing runs often rely on single-end reads to maximize turnaround time. We’ve overhauled our short read single-end assembly algorithms for WGS to produce significantly more contiguous assemblies, ensuring you don’t sacrifice genome quality for speed.
- Handling of unique sequencing data characteristics, such as two-color chemistry data and binned quality scores, the latter of which compresses the data for storage and transfers but reduces the granularity of data available to BugSeq.
Expanding AMR Databases for Precision Phenotype Prediction¶
Genomic antimicrobial susceptibility testing (AST) is revolutionizing how we understand and treat resistant infections. Building on our expert-augmented machine learning approach, we have expanded our curated AMR databases to enable even more accurate phenotype predictions for critical pathogens:
- General release of EUCAST-based AMR prediction: BugSeq now supports both EUCAST and CLSI breakpoints for the prediction of AMR. This expansion ensures greater accuracy and global applicability in determining resistance. To select EUCAST breakpoints, head to Analysis Configuration.
- Helicobacter pylori: H. pylori infects over half the world’s population, and rising antibiotic resistance is causing standard eradication therapies to fail at alarming rates (Tshibangu-Kabamba & Yamaoka, 2021). Because culturing H. pylori is notoriously difficult, Next-Generation Sequencing (NGS) is the ideal alternative. Our expanded database now accurately predicts H. pylori resistance profiles directly from sequence data.
- Mycobacterium tuberculosis vs. rifapentine: Rifapentine is an increasingly critical drug for both latent and active tuberculosis treatment regimens, particularly in shorter-course therapies that improve patient adherence. We have added rifapentine resistance markers to our TB database, allowing clinical and public health labs to monitor resistance and report genomic AST before phenotype is available.
- Meropenem-vaborbactam: Carbapenem-resistant Enterobacterales (CRE) are in the “critical group” of WHO’s bacterial priority pathogen list. Meropenem-vaborbactam is a new combination therapy designed to overcome certain beta-lactamases, such as KPC. Thanks to expert curation of our AMR database, BugSeq can now accurately predict susceptibility to this drug combination.
Conclusion¶
The first quarter of 2026 has set a strong foundation for the rest of the year. By enhancing our viral surveillance tools, optimizing for the latest rapid-sequencing hardware, deepening our AMR prediction capabilities, and building robust software bridges like our new EPI2ME integration, BugSeq continues to close the gap between complex genomic data and actionable clinical insights. We are committed to equipping our laboratory partners with the informatics needed to stay ahead of infectious diseases today and tomorrow.
Want to see a full list of changes? See our change log.
Have questions about how these updates impact your specific workflows? Reach out to our support team.